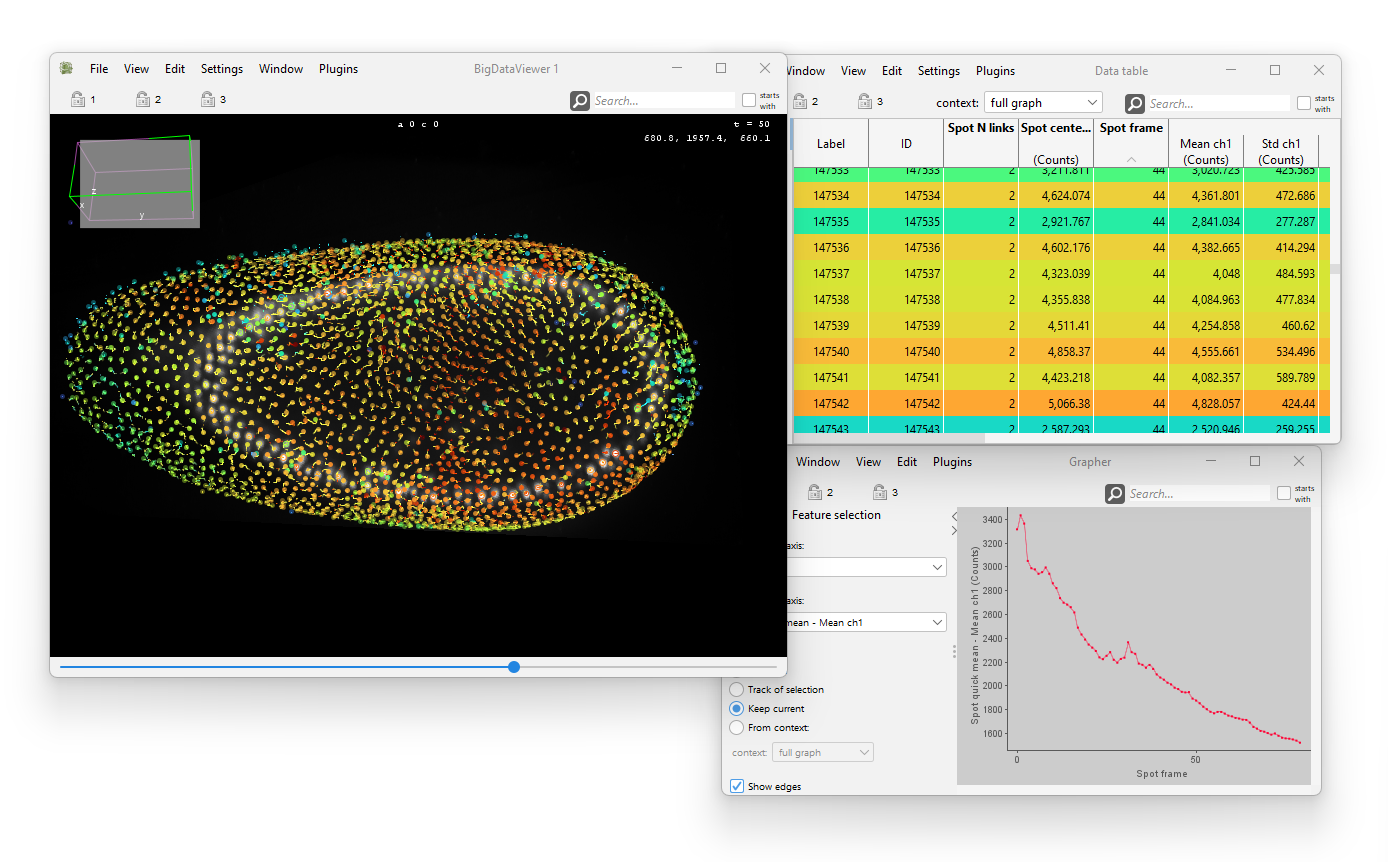
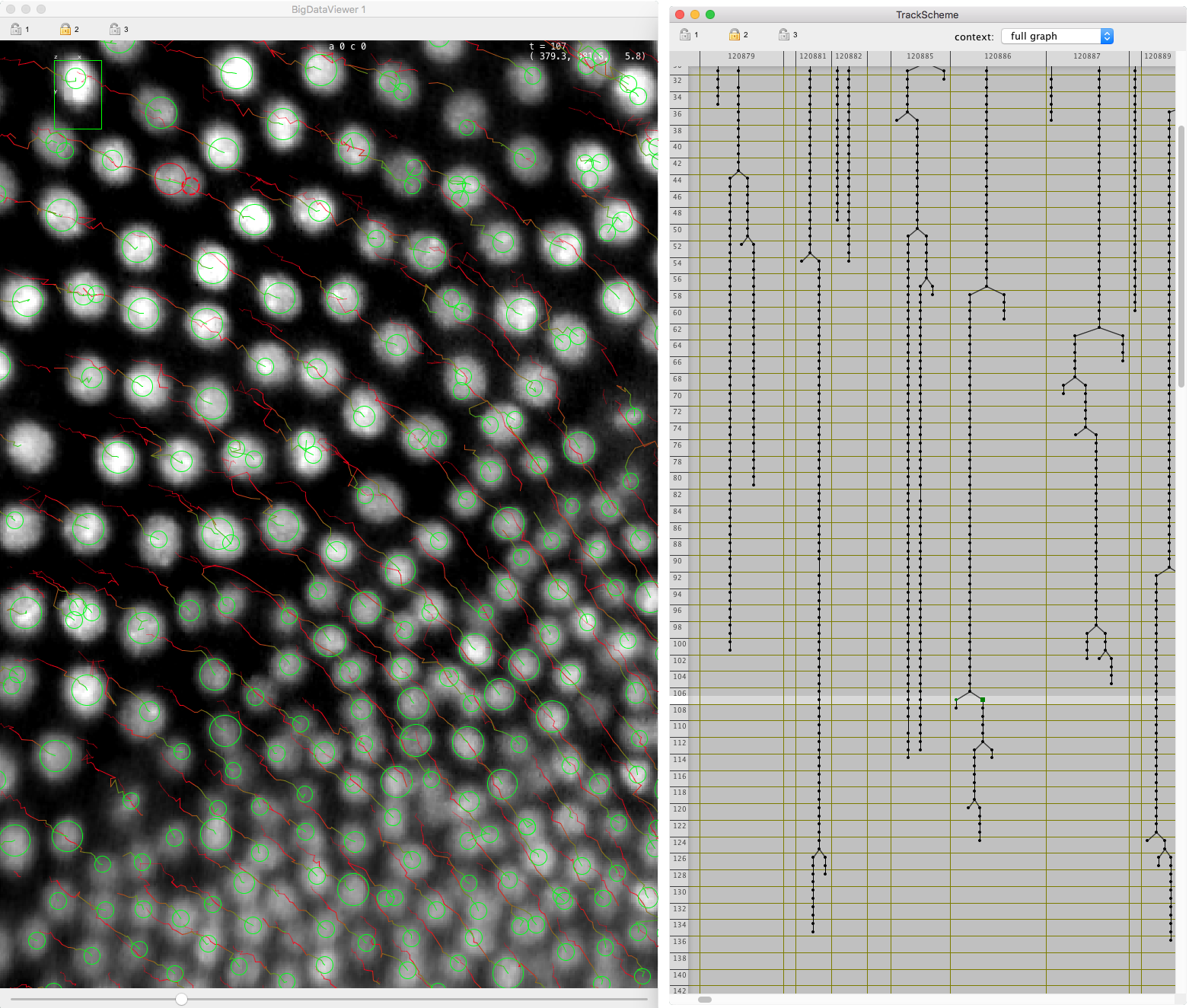
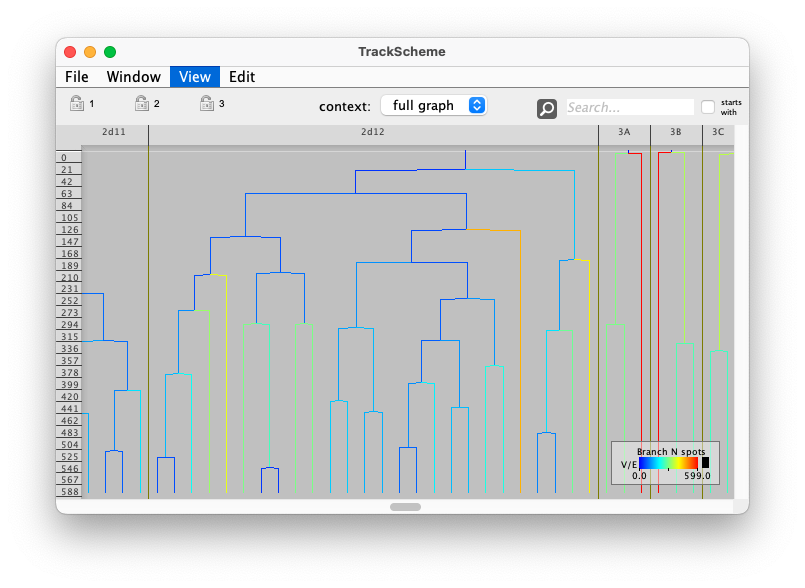
Mastodon is a large-scale tracking and track-editing framework for large, multi-view images, such as the ones that are typically generated in the domain Development Biology or Stem-Cell Biology or Cell Biology.

Why using Mastodon?
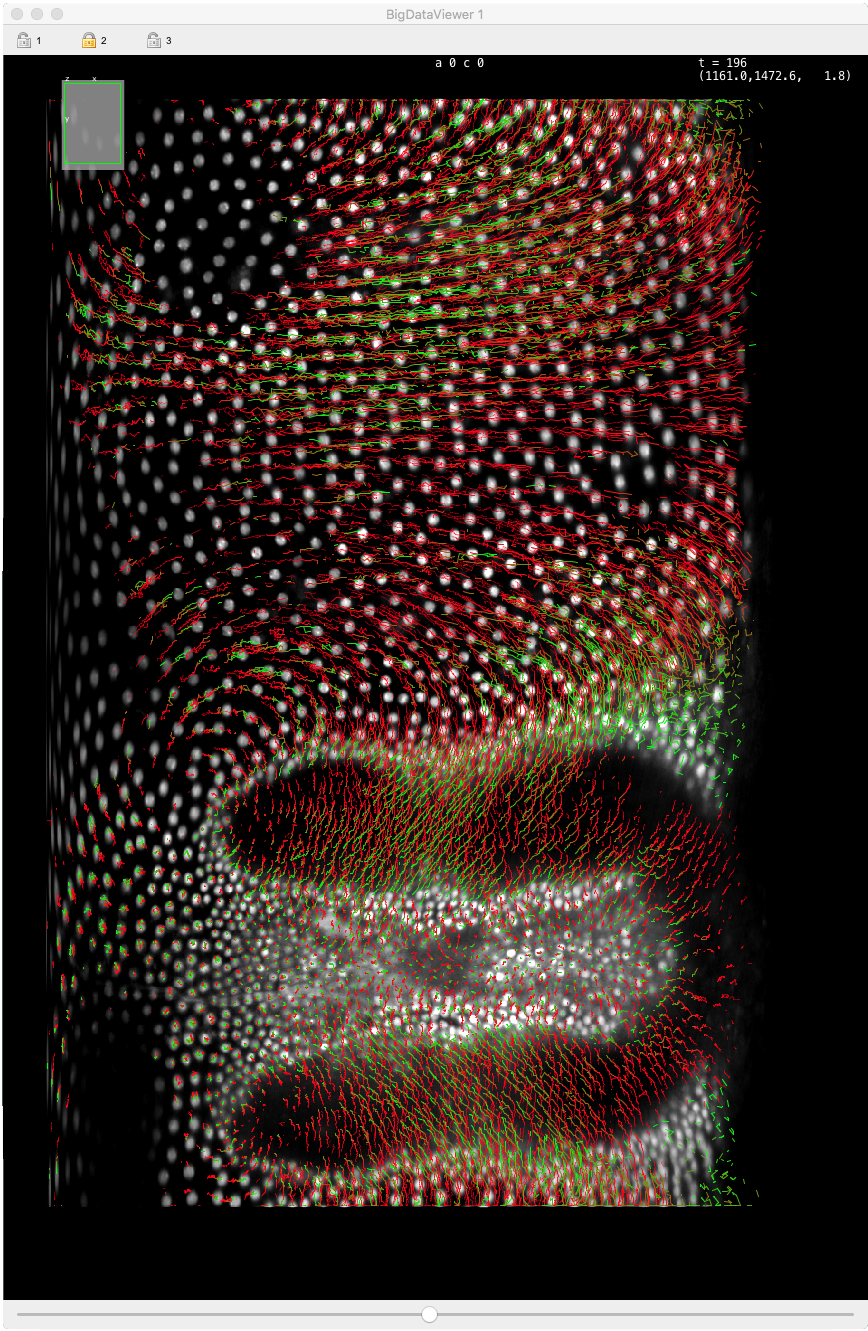
Modern microscopy technologies such as light sheet microscopy allows live sample in toto 3D imaging with high spatial and temporal resolution. Such images will be 3D over time, possibly multi-channels and multi-view. Computational analysis of these images promises new insights in cellular, developmental and stem cells biology. However, a single image can amount to several terabytes, and in turn, the automated or semi-automated analysis of these large images can generate a vast amount of annotations. The challenges of big data are then met twice: first by dealing with a very large image, and second with generating large annotations from this image. They will make interacting and analyzing the data especially difficult.
Mastodon is our effort to provide a tool that can harness these challenges.
You can find its documentation, tutorials and technical informations here.