The content of this page has not been vetted since shifting away from MediaWiki. If you’d like to help, check out the how to help guide!
This page supersedes the original Image5D page from the ImageJ documentation.
Please consider using the built-in hyperstacks functionality in ImageJ, since it is newer and more well-integrated than this old Image5D plugin.
Installation
Image5D is distributed as part of Fiji, so no installation is necessary beyond that. If you do not wish to enable the Fiji update site, you can download the Image5D JAR file to the plugins folder, restart ImageJ, and there will be new commands (New Image5D, Open Image5D, etc.) in the Plugins › Image5D submenu. Image5D requires ImageJ 1.37b or later.
Description
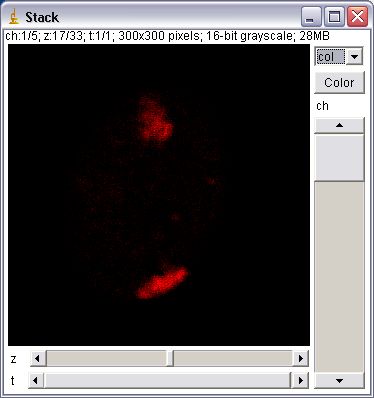
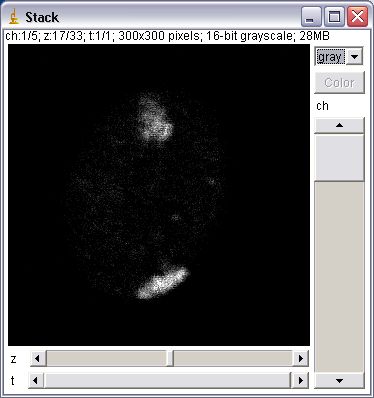
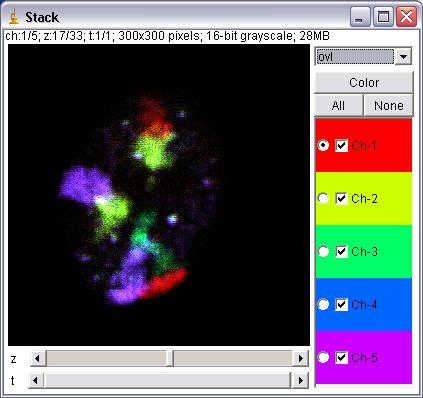
These plugins extend image stacks to 5 dimensions: x, y, channel (color), slice (z), frame (t). Unlike the HyperVolume Browser, an Image5D has a “true” 5D format. That way plugins working on stacks should usually work as expected on the currently displayed stack of an Image5D (just try a z-projection to see what this means). Image5Ds are displayed in a window with two scrollbars for slice and time below the image and a panel with controls to change the current channel and its color to the right of the image. A dropdown menu allows to change the display of the channels. Options are one channel in gray, one channel in color and an overlay of selected channels.
In the one channel color and overlay modes one can open a panel to select the display color of a channel or to edit a more refined LUT with the ImageJ LUT Editor.
To create an Image5D, use the plugins “Open Series as Image5D”, “Stack to Image5D” or “RGB to Image5D”.
Open Series as Image5D opens a series of images or stacks from hard disk and converts it to an Image5D. Stack to Image5D converts an already opened stack to an Image5D, and RGB to Image5D converts an RGB image or stack to an Image5D.
Missing features
- Saving just saves the current stack, not the full dataset.
- Calibration functions do not work.
- Converting the type (8-bit, 16-bit, float) does not work.
- Display changes in overlay mode are SLOW.
Known bugs
See the GitHub issue tracker
Sample data
A sample 5D data set is available. In consists of HeLa cells with an H2B-GFP construct to stain the chromatin imaged in fluorescence and transmission. A large part of the GFP fluorescence is bleached in the second frame and the cells go through mitosis, later. It has 2 channels, 24 slices and 7 frames. The file (Spindly-GFP.zip) is a TIFF stack contained in a ZIP archive. To view it with Image5D, open it using File › Open or drag and drop, then convert it to an Image5D using Plugins › Image5D › Stack to Image5D.
A sample 16-bit RGB data set of triple labeled cloned myoblast cells is also available. To view it with Image5D, open it using File › Open or drag and drop, then convert it to an Image5D using Plugins › Image5D › Stack to Image5D. This image is courtesy of Dr. Angelica Keller and Juliette Peltzer, Faculte des Sciences et Technologies, Universite Paris 12 Val de Marne, France.
See also
History
See the release notes for details on changes.