The content of this page has not been vetted since shifting away from MediaWiki. If you’d like to help, check out the how to help guide!
This plugin calculates the optic flow for each pair of images made with the given stack.
Principle
The plugin works using the PIV method, which is the most basic technique for optic flow. This technique, mainly used in acoustics or in fluids mechanics, enables the measurements of a velocity field in one plane, using imaging and image analysis1. It can be seen as one of the most simple pattern matching problem implementation.
PIV analysis is a block-based optic flow, based on inferring in what direction and in what amount a part of an image has moved between two successive instant. In the aforementioned domains, a flow is visualized by seeding it with light-reflecting particle (smoke in air, bubbles, glass beads in water, …) and imaged at two very close instants. The cross-correlation between parts of the two images where pattern generated by particles can be seen is then used to compute the velocity field.
The PIV algorithm is made of the following steps:
- two images are acquired of the same object are acquired at two successive instant;
- they are spliced in small pieces called interrogation windows;
- the cross-correlation between the two images is computed for each small window;
- the peak in the resulting correlation image is searched for. The peak location gives the displacement for which the two image parts look the best alike, that is: the amount by which the second image has to be moved to look like the first image the best;
- the velocity vector at this point is defined as the peak’s position. This assume that between the two successive instants, the image did not change too much in content, but moved or deformed.
Application to Life-science images
Here are two examples of its first applications in Biology:
- comparing flows in a drosophila embryo during gastrulation in control situations and after photo-ablation2;
- quantifying blood flow during cardiogenesis in zebrafish embryo3.
For us, the main interest of this technique is that it allows the computation of a velocity field without having to segment objects out of an image and track them, which makes it particularly interesting when dealing with brightfield or DIC images. The trade off is loss of precision, but also the fact that you might get completely irrelevant vectors.
Algorithm
As stated before, this plugin implements a very naive and primitive algorithm, without a sense of subtlety, and exists mainly for educational purpose. It has a very pedestrian approach, that make it slow. For each pixel away from the border of the image, a block is extracted for the front image and the back image. The correlation matrix is then calculated from these two blocks, and the result is analyzed to produce a flow vector. This is a highly redundant process, for when the algorithm moves to the next pixel to the right, the blocks content change only a little (only one column is replaced actually, and the rest is shifted left), but the whole correlation matrix is recalculated from scratch (a lot of wasted CPU cycles).
Typically, on a MacBook (grey model, 2009), for an 8-bit stack with a window size of 8x8, the plugin, the plugin can process a stack of 200x200 in approximately 1 second. I recommend downsampling the images, this would diminish the density of results, but make this plugin affordable.
The plugin is now multi-threaded, meaning that it will analyze multiple frame couples at once, depending how many CPUs it can find.
Example
The image to the right is made of 2000 small gaussian blobs (sigma=1), spawned randomly (you can get the original file here). The second frame was obtained by adding a shift to each blob, given by:
- ΔY = α × (X-Xm)
- ΔX = α × (Y-Ym)
with α=25 and Xm and Ym half of the image width. This effectively generate a sort of circular twist to the image.
Below is what the PIV analysis found, with a window of 64x64 (the max displacement the algorithm can find is given by half of the window size, so we had to go for at least 50x50), no masking and no interpolation. The color encode flow direction and magnitude, using the reference that can be seen in the color wheel.
Here are two other examples, taken from the MatPIV page of J. Kristian Sveen. The first one depict a surface wave in water:
The second one depict the internal wave. Notice the shearing in the middle of the frame.
Installation
Fiji
The plugin is part of the Fiji distribution. You can find it in the Plugins › Optic Flow menu.
Usage
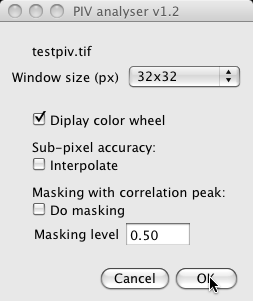
Options are the following:
- Window size: sets the interrogation window size for analysis. That is the size of the blocks that will be compared to get a flow out of it. See below for insight on size choices.
- Interpolate: if checked, the analysis will perform for each pixel an additional step, allowing to get sub-pixel flow magnitude. The maximum of the correlation window will be interpolated (using Taylor expansion over a 3x3 neighborhood), giving a better estimate of its position.
- Do masking: is set, results will be masked using the peak height value. For peak height values lower than Masking level ×
max(pkh), the flow will be set to 0.
ROI selection
If you want to restrain the analysis to only a certain portion of the image, select it in a ROI of any shape.
Image features
Tough you don’t need to have traceable objects for PIV, there is still some requirements for it.
PIV is based on a very simple pattern matching algorithm. This means that, to be effective, there must be some patterns in your image. Typically, the dotty structures of aggregate in a epi-fluorescence movie or the granularity that can be see on a brightfield image will do. As a rule of thumb if you can see something moving by eye, the PIV might be able to get it quantitatively; otherwise not of course.
Also keep in mind that (so far) it works only for a 2D plane. With optical sections, the algorithm might become puzzled by the appearance of grains moving in the Z direction. Also with thick specimen imaged in brightfield: the image of a dot deforms while moving in Z, which may lead to irrelevant values.
Choice of the interrogation window size
To perform pattern matching, the image is spliced in small pieces. Since you want to have clear patterns to match, it is a good idea to choose you interrogation window size bigger than the image feature whose movement you interested in.
Let us suppose that one one image, some structure can be found in the shape of ‘grains’ whose size is about 10-15 pixels. To always get at least a full one, a window of 32x32 pixels would be relevant.
Averaging
In fluid dynamics, velocity fields resulting from PIV are generally heavily spatially filtered afterward. In these cases, each of this vector represents velocity in a fluid for which the flow is generally spatially correlated on a large scale, so filtering in the spatial domain makes sense.
One can recommend to filter temporally, that is: for every vector in the field, replace its value by the moving average from a few frames before until a few frames after. If your movie is oversampled temporally, that might be effective in averaging outliers.
Filtering
The plugin returns also a stack made of the peak height for the analysis.
The peak height is just the value of the correlation peak for each pixels. This can be used to mask irrelevant part of the result image. The algorithm will be typically puzzled where there is no structure to correlate. In these parts of the image, the peak height will be typically low; one can then threshold the peak height image, and use it to mask the result image.
Version history
- 1.0 - 18 April 2009 - First public release.
- 1.1 - 19 April 2009 - can now be interrupted between frames.
- 1.2 - 20 April 2009 - is now multithreaded.