A Fiji plugin for the annotation of massive, multi-view data.
Installation
Enable the MaMuT update site in the Fiji updater to get MaMuT.
Publication
Presentation
MaMuT is an end user plugin that combines the BigDataViewer and TrackMate to provide an application that allow browsing, annotating and curating annotations for large image data.
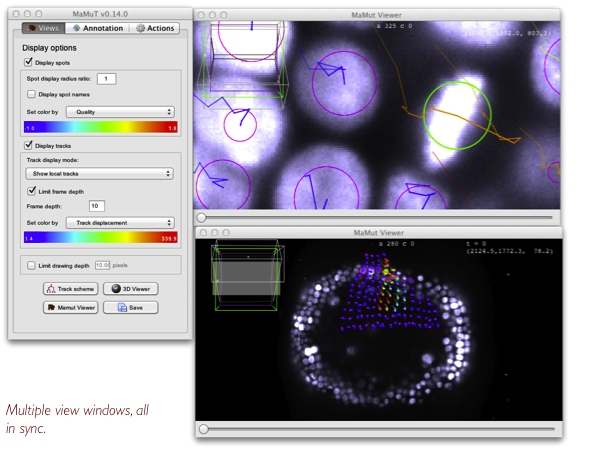
The main window resembles the display panel of TrackMate. It controls how the annotations are displayed. Using the MaMuT Viewer button, several views of the data can be launched. They will all be in sync. Each of them is an instance of the BigDataViewer.
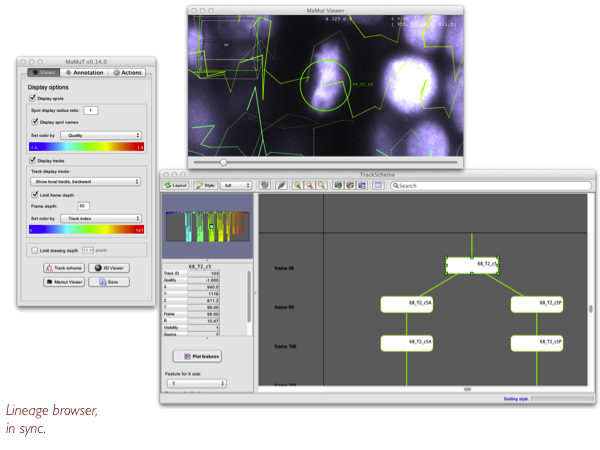
We privileged annotations that are like lineages, or object followed over time (which is what TrackMate does). MaMuT ships TrackScheme, the lineage browser taken from TrackMate.
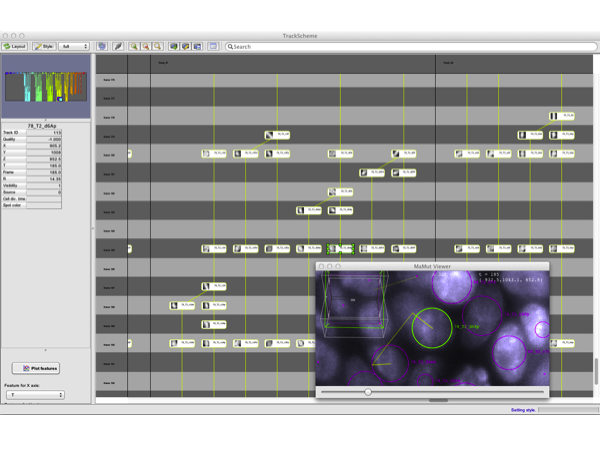
However, MaMuT itself does not ship any fully-automated or tracking algorithm. It is meant for manual or semi-automatic annotation. Still, we made the GUI comfortable enough so that you can quickly generate rather large annotations. A semi-automated segmentation can help you generating quickly lineage branches from single cells.
User documentation
The following pages are tutorials that will guide you through MaMuT and walk you though its features.
- Getting started with MaMuT is an introduction tutorial. It will show you how to prepare a dataset for MaMuT and make your first annotations with it.
- Example mamut.properties file contains an example of a
mamut.propertiesfile, that is used to customize the key-bindings in the MaMuT viewer.