The content of this page has not been vetted since shifting away from MediaWiki. If you’d like to help, check out the how to help guide!
The default algorithm to identify the seeds as objects is called Maximally Stable Extremal Regions (MSER). In short, it computes all possible intensity thresholds of an image, which is called a component tree (e.g. it creates a 3d image from a 2d image). Subsequently MSER tries to identify “stable regions” in this space. Regions in the component tree are considered stable if an object has a certain size and preserves its shape over a given range of thresholds without merging with other objects.
MSER detected regions as shown as red ellipses. The line along which each of the Microtubules lie is assumed to be the semi-major axis of the MSER ellipse. The MSER is computed on an unsigned byte image where intensity levels are integers between 0-255 and several parameters can be adjusted to find as many microtubules as possible:
- Threshold difference defines the resolution of the component tree, i.e. every how many intensity values a threshold is computed. A low value should be used when low intensity objects are to be recognized. Increasing the parameter value will only allow higher intensity objects to be recognized.
- Stability Score: is a threshold for the minimal stability of a region in the component space. For values close to 1 many regions will be deemed stable and for values close to 0 only very few extremely stable regions will be found.
- Min # of pixels: This number vetoes the regions which have fewer pixels than this pre-set value and includes only the ellipses which carry more pixels than this value.
- Max # of pixels (Advanced Mode): This number vetoes the regions which have more pixels than this pre-set value and only includes the ellipses which carry fewer pixels than this value. It is only available in the advanced mode and in the simple mode it is pre-set to image dimension in X times the image dimension in Y.
- Min diversity (Advanced Mode): The MSER algorithm creates a component tree and based on this parameter user can choose to keep all the regions or to exclude some of the regions. For values equal to 1 only the roots of the component trees would be kept as stable regions, by decreasing this value more MSER regions would show starting from regions which are closest to the roots in the component tree and for small values of the parameter even the regions which are farthest away from the roots of the component tree would show up. In the simple mode the value is kept at 1 and in the advanced mode this value can be chosen as per user discretion. For seed images a value of 1 is recommended. During tracking however, this parameter is helpful in avoiding miss-assignments by the optimizer if the microtubules collide for example, values above 0.75 are recommended in such cases.
As an example we show in the animation below how during a collision including the children of the component tree makes sub-elliptical regions inside the main MSER ellipse, when the assignment of the end points is done it is more likely to do a proper assignment during such a collision when the value of min diversity was set to a value less than unity (0.8 in this case), on the right side of the animation this parameter was unity and was more likely to make a mistake in doing the assignment of the end points (was avoided due to good initial guesses given to the optimizer).

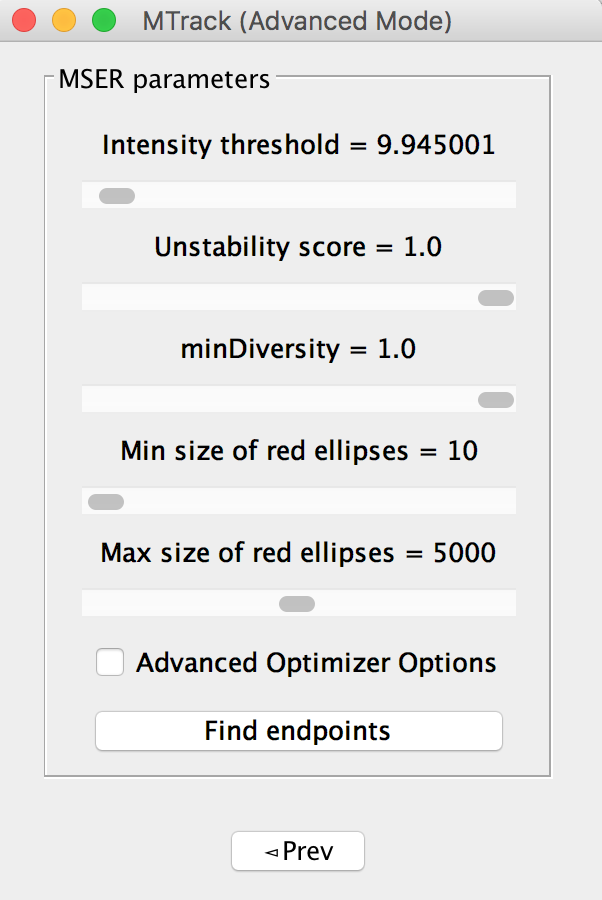
MSER panel in the advanced mode appears as shown below, after clicking on “find endpoints” a next button would appear along with the prev button, allowing user to flip to the next panel.